Andrea Wilderman
Graduate Student
WILDERMAN_ANDREA_OCTOBER_2021_CV
Full Bibliography
https://www.ncbi.nlm.nih.gov/myncbi/andrea.wilderman.1/bibliography/public/
Computational Prediction and Experimental Validation of Tissue-Specific Superenhancers
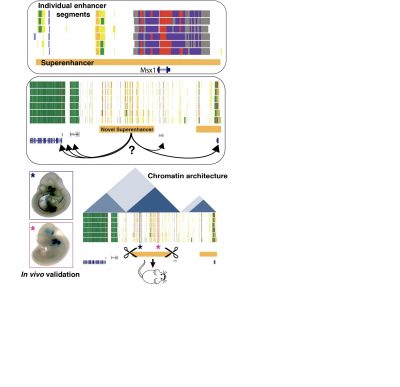
The regulatory programs that build and shape the face require precisely controlled spatiotemporal gene expression, achieved through tissue-specific enhancers. Groups of coordinately activated enhancers, often referred to as ‘superenhancers’, encompass and regulate genes important in determining cell identity. My work uses both computational and experimental methods to identify tissue-specific superenhancer regions and predict, then confirm the genes they regulate. Our lab has generated epigenomic atlases of human embryonic heart and craniofacial tissue and identified superenhancer regions specific to each tissue. My thesis project centers on a craniofacial-specific superenhancer in a large gene desert on human Chromosome 7. The neighboring genes are not known to have relevance to craniofacial development. I have used the chromatin conformation capture method HiC to identify potential targets of this superenhancer region and confirmed long-range interactions between the superenhancer CV and predicted target using a higher-resolution chromatin conformation capture method, 4C-seq. Deletion of this region in mouse creates a restructuring of the local chromatin architecture, yet produces a very limited effect on gene expression. Mice homozygous for deletion of the superenhancer confirm the tissue-specificity of the enhancers, demonstrate that the region is essential for viability and have high penetrance of orofacial clefts. This evidence suggests we have identified a critical non-coding regulatory region with highly craniofacial-specific effects whose deletion could be pathogenic in human patients.
Examples of HiC data processing pipeline
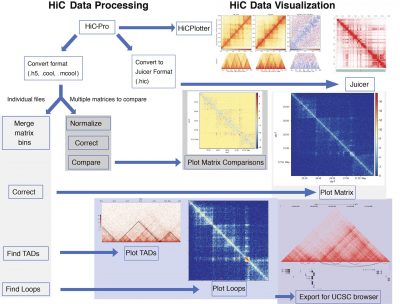
Learn how:
https://github.com/awilderman/HiC

| awilderman@uchc.edu | |
| Phone | 860-679-5597 |
| Office Location | R1261, 400 Farmington Ave. |
| Campus | Uconn Health |